More from All
APIs in general are so powerful.
Best 5 public APIs you can use to build your next project:
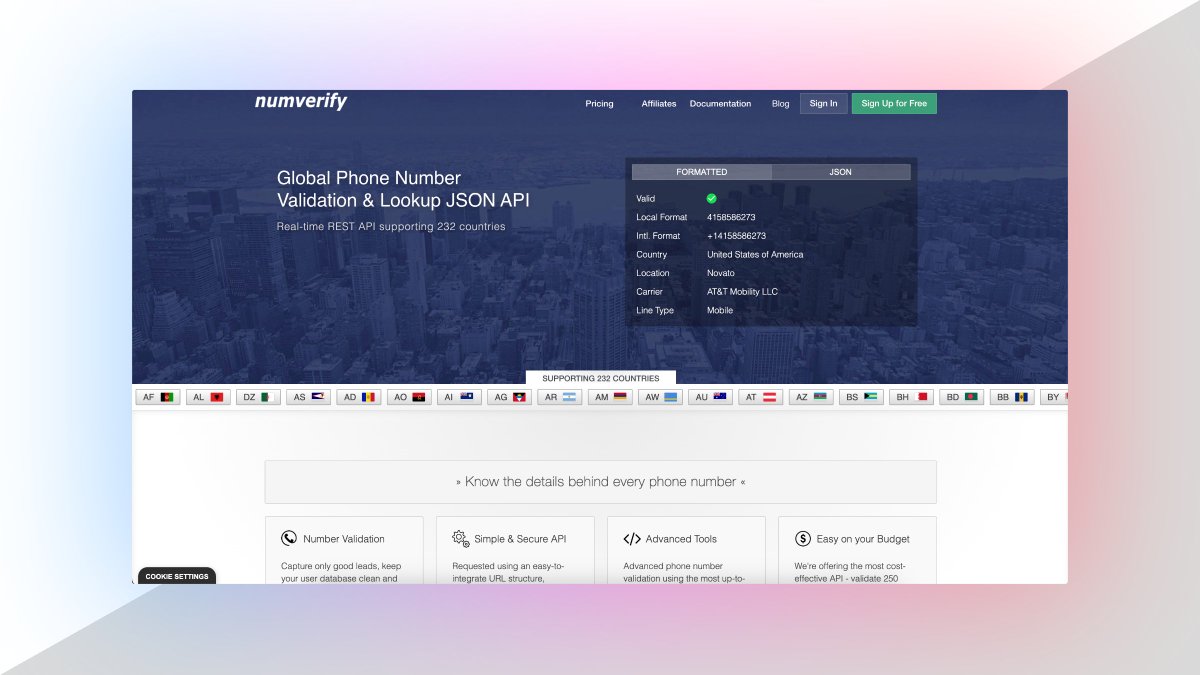
1. Number Verification API
A RESTful JSON API for national and international phone number validation.
🔗 https://t.co/fzBmCMFdIj
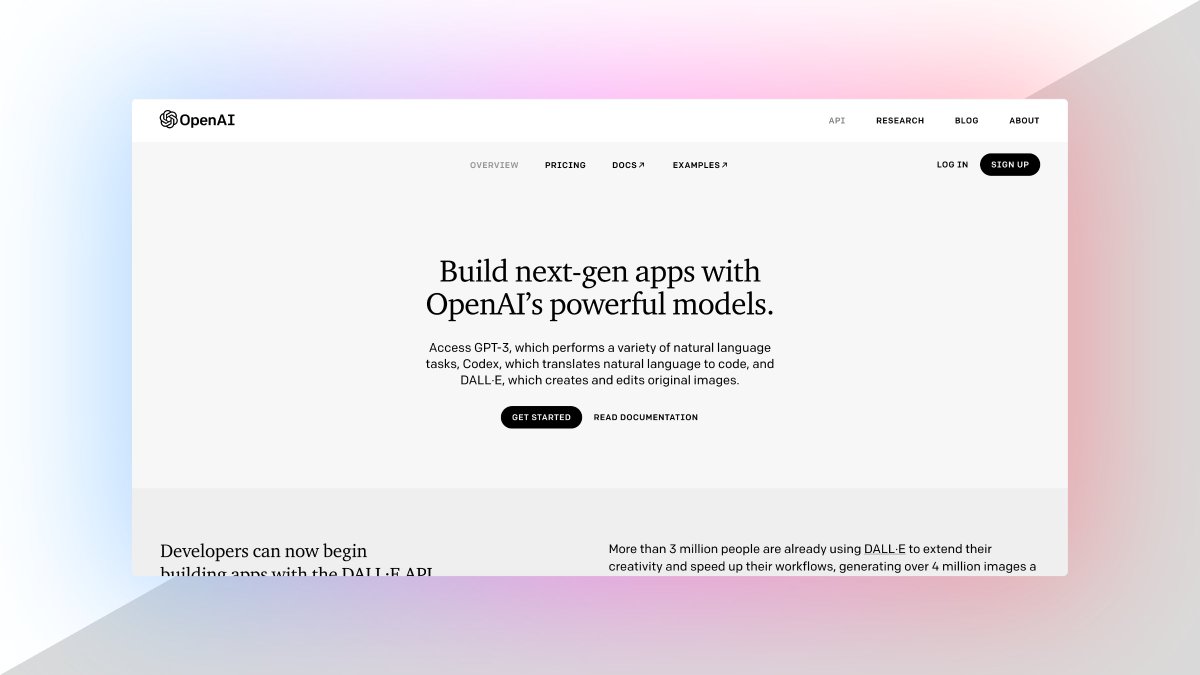
2. OpenAI API
ChatGPT is an outstanding tool. Build your own API applications with OpenAI API.
🔗 https://t.co/TVnTciMpML
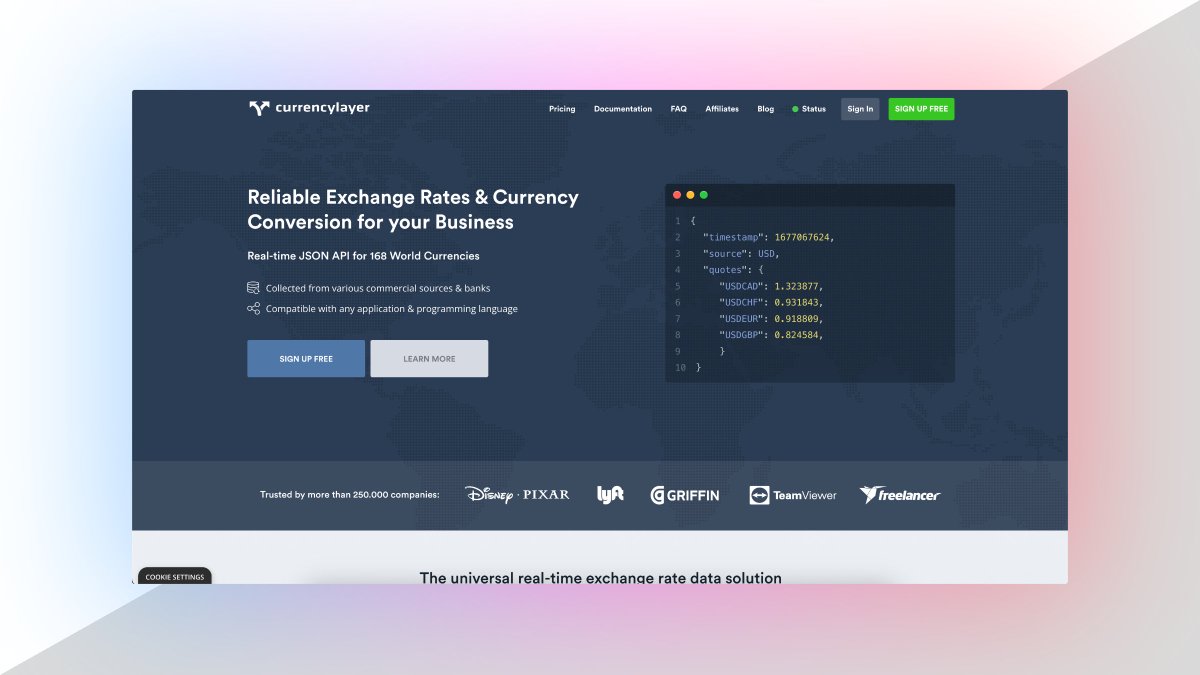
3. Currency Data API
Currency Data API provides a simple REST API with real-time and historical exchange rates for 168 world currencies
🔗 https://t.co/TRj35IUUec
4. Weather API
Real-Time & historical world weather data API.
Retrieve instant, accurate weather information for
any location in the world in lightweight JSON format.
🔗 https://t.co/DCY8kXqVIK
Best 5 public APIs you can use to build your next project:
1. Number Verification API
A RESTful JSON API for national and international phone number validation.
🔗 https://t.co/fzBmCMFdIj

2. OpenAI API
ChatGPT is an outstanding tool. Build your own API applications with OpenAI API.
🔗 https://t.co/TVnTciMpML

3. Currency Data API
Currency Data API provides a simple REST API with real-time and historical exchange rates for 168 world currencies
🔗 https://t.co/TRj35IUUec

4. Weather API
Real-Time & historical world weather data API.
Retrieve instant, accurate weather information for
any location in the world in lightweight JSON format.
🔗 https://t.co/DCY8kXqVIK