1. The same “gang” of the Proximal Origin paper
https://t.co/Xmp20I58AQ
in action again (only Lipkin is missing, who knows why) to push once more faulty arguments in support of SARS2’s natural
https://t.co/0Goapx6cXj
https://t.co/5Z4Ndca98t
https://t.co/i9OZchD4LT
“adjusted by visual inspection” to push their false conclusion that all these CoVs have a partial FCS insertion at the S1/S2 junction and the FCS of SARS2 is therefore natural
https://t.co/viqvfzv1Xa
and that can be easily inserted with the Seamless technology.
https://t.co/U2qvG3tXBG
https://t.co/cNEwlgtnBb
https://t.co/1D7iVZc8Md
9/ Moreover, the QTQTN motive proximal to the FCS is beneficial for virus entry in presence of Cathepsin, which is naturally produced by kidney cells. https://t.co/xPOK6w931P
— Rossana Segreto (@Rossana38510044) October 3, 2020
https://t.co/GFrrxzCvbq
https://t.co/pBF4ZDqNat
You know virology is broken when one top virologist approvingly retweets another top virologist's complete nonsense.
— Yuri Deigin (@ydeigin) February 23, 2021
Look at the QTQTNS fragment's underlying nucleotides - they are identical. No way this has "arisen independently in multiple bat sarbecoviruses". EvoBio 101 FAIL! pic.twitter.com/SNxesiwYPg
https://t.co/DjbJm40hni
https://t.co/lp9C8vtBWg
https://t.co/viqvfzv1Xa
And of course, after millions of passages in humans the virus can mutate to bind even better than possibly obtained in cell culture or humanized mouse.
More from All
You May Also Like
Keep dwelling on this:
Further Examination of the Motif near PRRA Reveals Close Structural Similarity to the SEB Superantigen as well as Sequence Similarities to Neurotoxins and a Viral SAg.
The insertion PRRA together with 7 sequentially preceding residues & succeeding R685 (conserved in β-CoVs) form a motif, Y674QTQTNSPRRAR685, homologous to those of neurotoxins from Ophiophagus (cobra) and Bungarus genera, as well as neurotoxin-like regions from three RABV strains
(20) (Fig. 2D). We further noticed that the same segment bears close similarity to the HIV-1 glycoprotein gp120 SAg motif F164 to V174.
https://t.co/EwwJOSa8RK
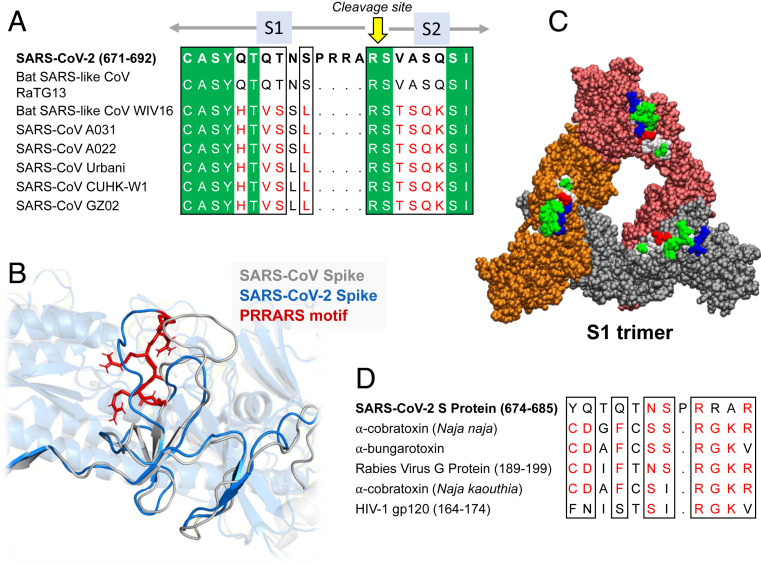
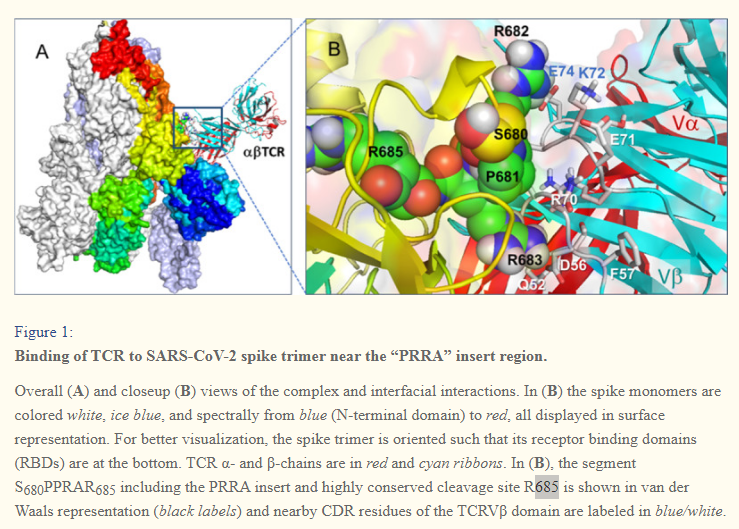
In (B), the segment S680PPRAR685 including the PRRA insert and highly conserved cleavage site *R685* is shown in van der Waals representation (black labels) and nearby CDR residues of the TCRVβ domain are labeled in blue/white
https://t.co/BsY8BAIzDa
Sequence Identity %
https://t.co/BsY8BAIzDa
Y674 - QTQTNSPRRA - R685
Similar to neurotoxins from Ophiophagus (cobra) & Bungarus genera & neurotoxin-like regions from three RABV strains
T678 - NSPRRA- R685
Superantigenic core, consistently aligned against bacterial or viral SAgs
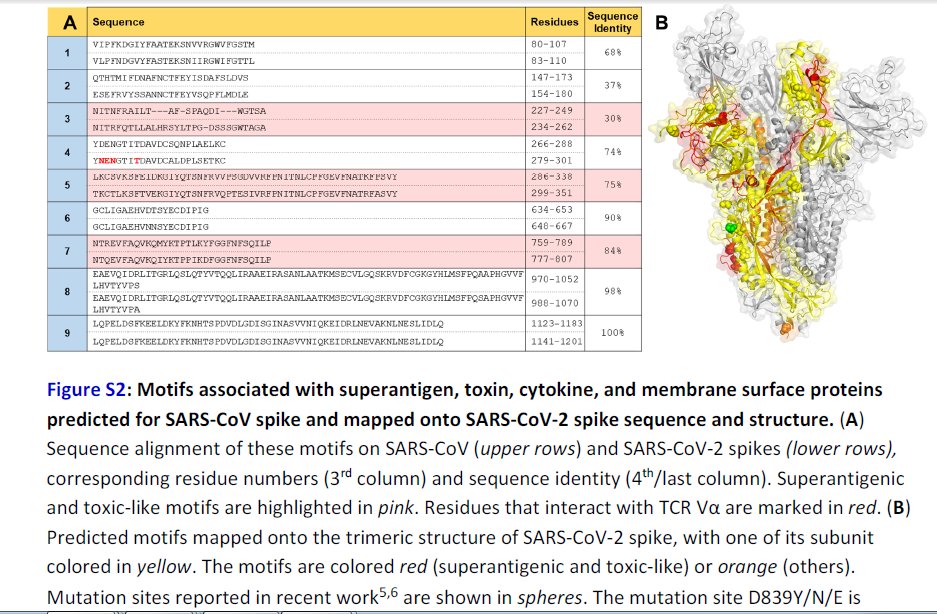

Further Examination of the Motif near PRRA Reveals Close Structural Similarity to the SEB Superantigen as well as Sequence Similarities to Neurotoxins and a Viral SAg.
The insertion PRRA together with 7 sequentially preceding residues & succeeding R685 (conserved in β-CoVs) form a motif, Y674QTQTNSPRRAR685, homologous to those of neurotoxins from Ophiophagus (cobra) and Bungarus genera, as well as neurotoxin-like regions from three RABV strains
(20) (Fig. 2D). We further noticed that the same segment bears close similarity to the HIV-1 glycoprotein gp120 SAg motif F164 to V174.
https://t.co/EwwJOSa8RK

In (B), the segment S680PPRAR685 including the PRRA insert and highly conserved cleavage site *R685* is shown in van der Waals representation (black labels) and nearby CDR residues of the TCRVβ domain are labeled in blue/white
https://t.co/BsY8BAIzDa

Sequence Identity %
https://t.co/BsY8BAIzDa
Y674 - QTQTNSPRRA - R685
Similar to neurotoxins from Ophiophagus (cobra) & Bungarus genera & neurotoxin-like regions from three RABV strains
T678 - NSPRRA- R685
Superantigenic core, consistently aligned against bacterial or viral SAgs