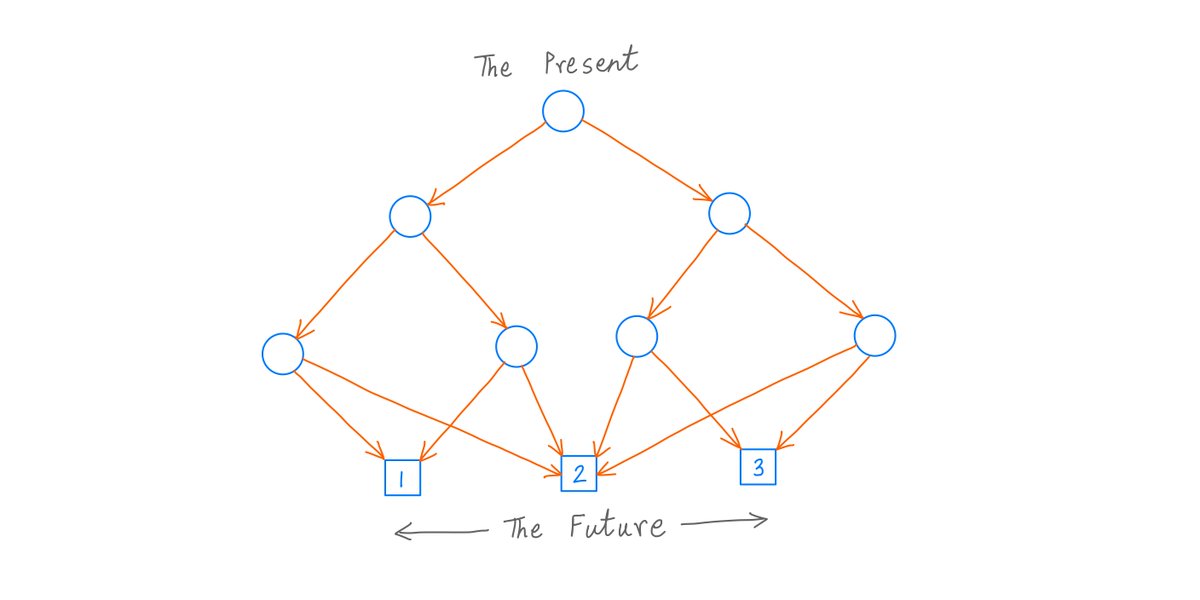
Recently I learned something about DNA that blew my mind, and in this thread, I'll attempt to blow your mind as well. Behold: Chargaff's 2nd Parity Rule for DNA N-Grams.
If you are into cryptography or reverse engineering, you should love this.
Thread:


T G T C A G T
A C A G T C A
(note how the other strand is upside down - this matters!)

This is called Chargaff's 1st parity rule.
https://t.co/jD4cMt0PJ0

This is called Chargaff's 2nd parity rule.


For N=2, this says that percentage of CC (%CC) and %GG are also equal, as are %AG and %CT (complemented AND reversed) etc.



More from Science
1. I find it remarkable that some medics and scientists aren’t raising their voices to make children as safe as possible. The comment about children being less infectious than adults is unsupported by evidence.
2. @c_drosten has talked about this extensively and @dgurdasani1 and @DrZoeHyde have repeatedly pointed out flaws in the studies which have purported to show this. Now for the other assertion: children are very rarely ill with COVID19.
3. Children seem to suffer less with acute illness, but we have no idea of the long-term impact of infection. We do know #LongCovid affects some children. @LongCovidKids now speaks for 1,500 children struggling with a wide range of long-term symptoms.
4. 1,500 children whose parents found a small campaign group. How many more are out there? We don’t know. ONS data suggests there might be many, but the issue hasn’t been studied sufficiently well or long enough for a definitive answer.
5. Some people have talked about #COVID19 being this generation’s Polio. According to US CDC, Polio resulted in inapparent infection in more than 99% of people. Severe disease occurred in a tiny fraction of those infected. Source:
I find it remarkable that a section of society not rejoicing that children very rarely ill with COVID compared to other viruses and much less infectious than adults
— Michael Absoud \U0001f499 (@MAbsoud) February 12, 2021
Instead trying prove the opposite!
Why??
2. @c_drosten has talked about this extensively and @dgurdasani1 and @DrZoeHyde have repeatedly pointed out flaws in the studies which have purported to show this. Now for the other assertion: children are very rarely ill with COVID19.
3. Children seem to suffer less with acute illness, but we have no idea of the long-term impact of infection. We do know #LongCovid affects some children. @LongCovidKids now speaks for 1,500 children struggling with a wide range of long-term symptoms.
4. 1,500 children whose parents found a small campaign group. How many more are out there? We don’t know. ONS data suggests there might be many, but the issue hasn’t been studied sufficiently well or long enough for a definitive answer.
5. Some people have talked about #COVID19 being this generation’s Polio. According to US CDC, Polio resulted in inapparent infection in more than 99% of people. Severe disease occurred in a tiny fraction of those infected. Source:
You May Also Like
#sculpture #story -
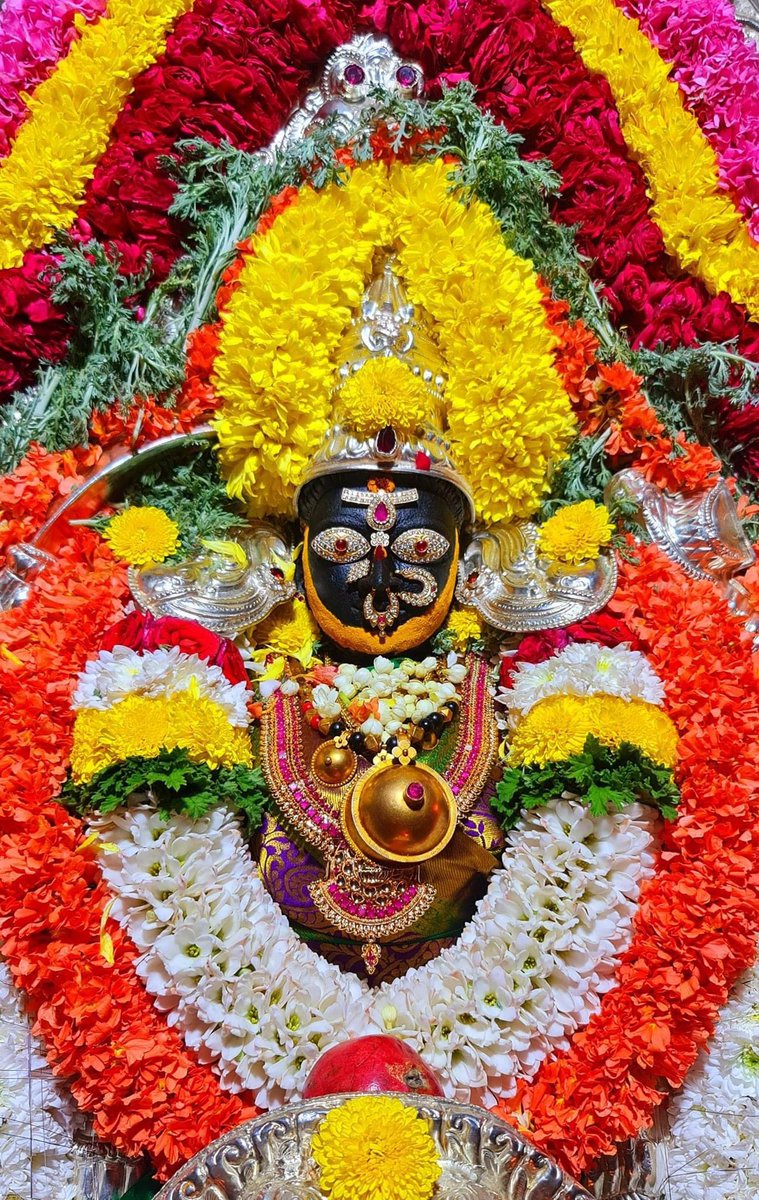
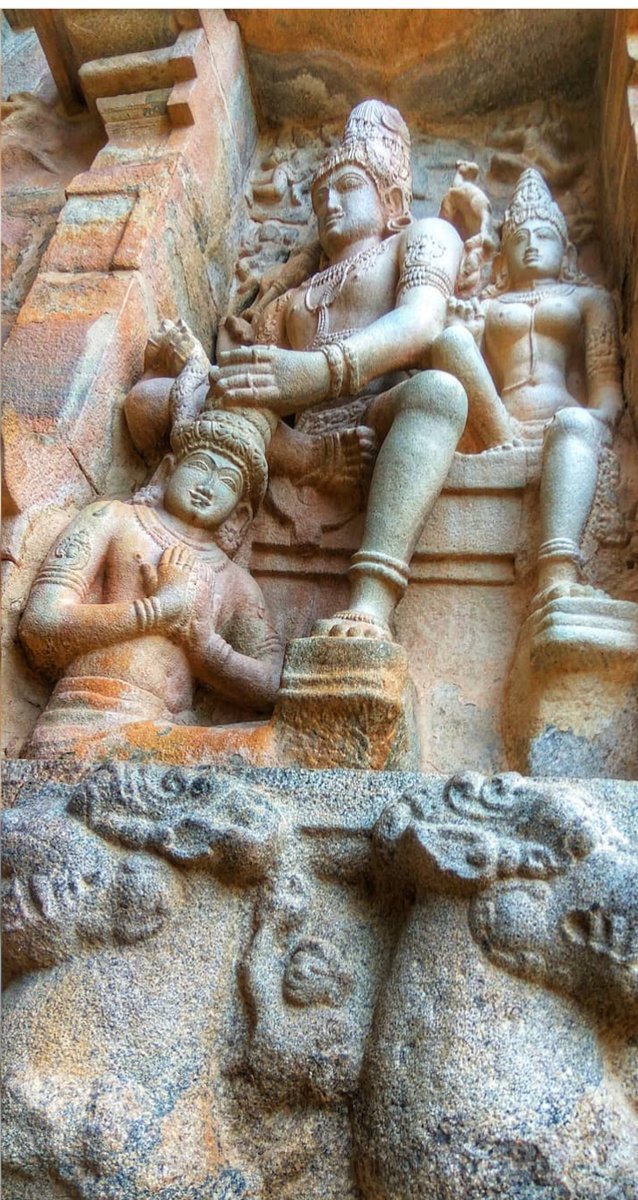
Chandesha-Anugraha Murti - One of the Sculpture in Brihadeshwara Temple at Gangaikonda Cholapuram - built by Raja Rajendra Chola I
This Sculpture depicts Bhagwan Shiva along with Devi Paravathi blessing Chandeshwara - one of the 63 Nayanmars.
#Thread

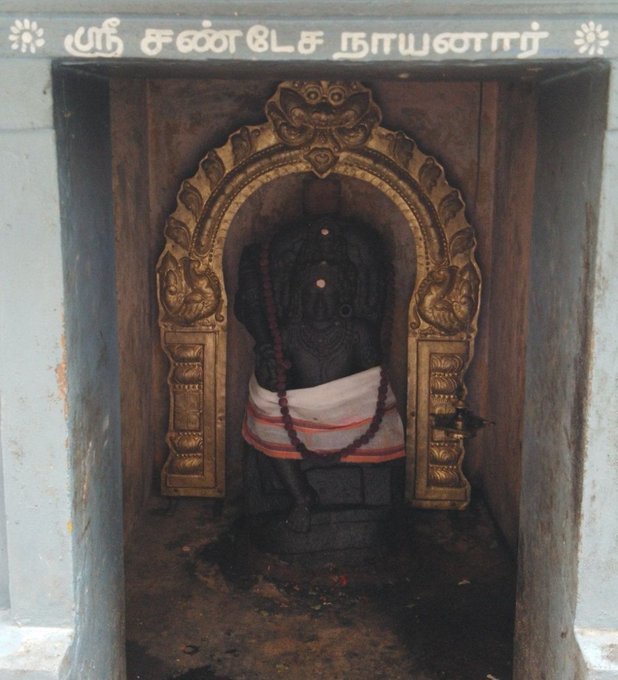
Chandeshwara/Chandikeshwara is regarded as custodian of Shiva Temple's wealth&most of Shiva temples in South India has separate sannathi for him.
His bhakti for Bhagwan Shiva elevated him as one of foremost among Nayanmars.
He gave importance to Shiva Pooja&protection of cows.

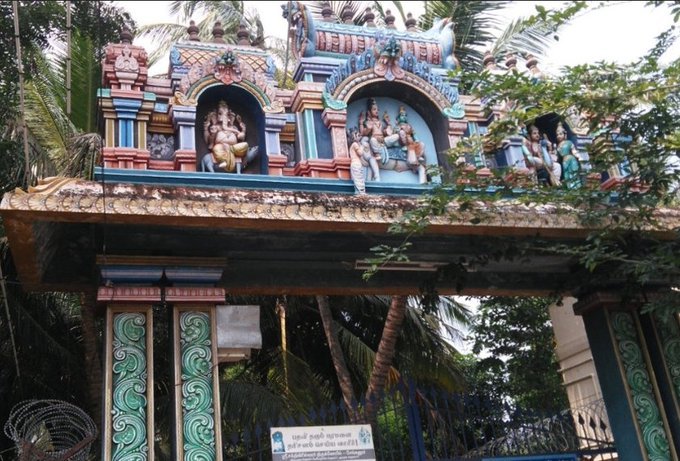
There are series of paintings, illustrating the #story of Chandikeshwar in the premises of
Sri Sathiyagireeswarar #Temple at Seinganur,near Kumbakonam,TN
Chandikeshwara's birth name
is Vichara sarman.He was born in the village of Senganur on the banks of River Manni.

His Parent names were Yajnathatan and Pavithrai.
Vichara Sarman was a gifted child and he learnt Vedas and Agamas at a very young age.
He was very devout and would always think about Bhagwan Shiva.
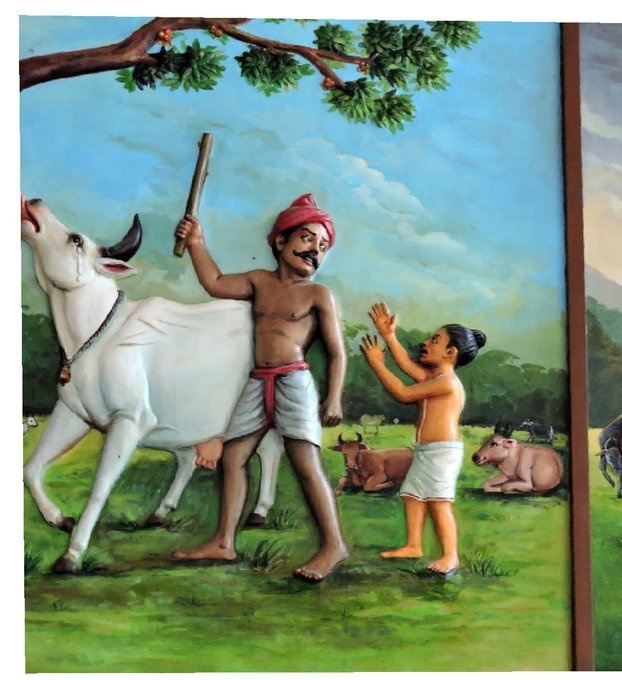
One day he saw a cowherd man brutally assaulting a cow,Vichara Sarman could not tolerate this. He spoke to cowherd: ‘Do you not know that the cow is worshipful & divine? All gods & Devas reside in https://t.co/ElLcI5ppsK it is our duty to protect cows &we should not to harm them.

Chandesha-Anugraha Murti - One of the Sculpture in Brihadeshwara Temple at Gangaikonda Cholapuram - built by Raja Rajendra Chola I
This Sculpture depicts Bhagwan Shiva along with Devi Paravathi blessing Chandeshwara - one of the 63 Nayanmars.
#Thread

Chandeshwara/Chandikeshwara is regarded as custodian of Shiva Temple's wealth&most of Shiva temples in South India has separate sannathi for him.
His bhakti for Bhagwan Shiva elevated him as one of foremost among Nayanmars.
He gave importance to Shiva Pooja&protection of cows.

There are series of paintings, illustrating the #story of Chandikeshwar in the premises of
Sri Sathiyagireeswarar #Temple at Seinganur,near Kumbakonam,TN
Chandikeshwara's birth name
is Vichara sarman.He was born in the village of Senganur on the banks of River Manni.

His Parent names were Yajnathatan and Pavithrai.
Vichara Sarman was a gifted child and he learnt Vedas and Agamas at a very young age.
He was very devout and would always think about Bhagwan Shiva.
One day he saw a cowherd man brutally assaulting a cow,Vichara Sarman could not tolerate this. He spoke to cowherd: ‘Do you not know that the cow is worshipful & divine? All gods & Devas reside in https://t.co/ElLcI5ppsK it is our duty to protect cows &we should not to harm them.